Optimization of parameters or “systems” in general plays an ever-increasing role in mathematics, economics, engineering, and the life sciences. As a result, a wide variety of both traditional mathematical and nontraditional algorithmic approaches have been introduced to solve challenging and practically relevant optimization problems. Evolutionary optimization methods—in the form of genetic algorithms, genetic programming, and evolution strategies—represent nontraditional optimization algorithms that draw inspiration from the processes of natural evolution. Particle swarm optimization is another set of more recently developed algorithmic optimizers inspired by social behaviors of organisms such as birds [1] and social insects. These new evolutionary approaches in optimization are now entering the stage and are becoming very successful tools for solving real-world optimization problems [2]. We present Visplore and Evolvica as a toolkit to investigate, explore, and visualize evolutionary and swarm-based optimization techniques. A webMathematica interface is also available.
1. Introduction
The evolutionary optimization methods of the genetic algorithm (GA) [3], genetic programming (GP) [4], and evolution strategy (ES) [5] are a branch of nontraditional optimization methods drawing inspiration from the processes of natural evolution. The particle swarm optimizer (PSO), on the other hand, is inspired by the social behavior of bird flocking [6]. Recently, we have been investigating the performance of evolution- and swarm-based optimizers in the domain of biomechanics, which we developed with the Human Performance Laboratory at the Faculty of Kinesiology, University of Calgary [7–9]. In this particular biomechanical application, numerical optimization algorithms are used to design equipment for sports activities. The involved simulations of muscle movements are very time consuming and high dimensional, thus making their evaluation costly and difficult. Simulating a soccer kick is an example of a model that investigates muscle activation patterns within the leg and foot when kicking a soccer ball toward the goal. The specific objective in this case is to obtain a high ball speed, in order to minimize the goal keeper’s chances of catching the ball. In 1998, Cole applied a ![]() ES to this model [7, 8]. More recently, we presented improved adaptations of the model parameters through a PSO [2, 10].
ES to this model [7, 8]. More recently, we presented improved adaptations of the model parameters through a PSO [2, 10].
The main focus of this article, however, is not on the optimization of parameters for the soccer kick model. Instead, we present what has been learned from our comparison studies of the evolution- and swarm-based optimizers on a set of selected benchmark functions. These benchmark studies turned out to be extremely useful in understanding the intricacies in performance regarding three optimizers: (1) the originally used ![]() ES; (2) a canonical (“basic”) PSO (bPSO); and (3) a PSO with noise-induced (“random”) inertia weight settings (rPSO). We describe and analyze the performance of each of these optimizers on five benchmark functions in two, four, and 10 dimensions. These findings are projected to performance characteristics that were found in the real-world application of the discussed soccer-kick model, which poses a 56-dimensional optimization problem. The Mathematica notebooks that were created provide us with insights regarding the relations between control parameters and system performance of the optimizers under study. Consequently, we gain a better understanding of the algorithms on multidimensional real-world problems.
ES; (2) a canonical (“basic”) PSO (bPSO); and (3) a PSO with noise-induced (“random”) inertia weight settings (rPSO). We describe and analyze the performance of each of these optimizers on five benchmark functions in two, four, and 10 dimensions. These findings are projected to performance characteristics that were found in the real-world application of the discussed soccer-kick model, which poses a 56-dimensional optimization problem. The Mathematica notebooks that were created provide us with insights regarding the relations between control parameters and system performance of the optimizers under study. Consequently, we gain a better understanding of the algorithms on multidimensional real-world problems.
This article is organized as follows. In Section 2 we give descriptions of the three optimization algorithms used in our comparison. An introduction to the benchmark functions and an outline of the experimental setup follows in Sections 3 and 4, respectively. We discuss the experimental results and summarize the lessons learned in Section 5. The accompanying ![]() site is presented in Section 6. The paper is concluded in Section 7.
site is presented in Section 6. The paper is concluded in Section 7.
2. The Three Contenders
The three contenders for our comparative study of evolution- and swarm-based optimization algorithms are: (1) a relatively simple ![]() ES; (2) a canonical (“basic”) PSO (bPSO); and (3) a PSO with noise-induced (“random”) inertia weight settings (rPSO). The following subsections present these approaches in more detail.
ES; (2) a canonical (“basic”) PSO (bPSO); and (3) a PSO with noise-induced (“random”) inertia weight settings (rPSO). The following subsections present these approaches in more detail.
2.1.  Evolution Strategy
Evolution Strategy
ES has been a successful evolutionary technique for solving complex optimization problems since the 1960s [5]. ES evolves vectors of real numbers and the “genetic” information is interchanged between these vectors through recombination operators. Slight variations (“mutations”) on these vectors are obtained by evolving strategy parameters that determine the amount of change applied to each vector component.
In the ![]() ES scheme, a single parent is mutated
ES scheme, a single parent is mutated ![]() times. Each of the newly created offspring is evaluated, and the parents and the offspring are added to the selection pool. The single best individual among the
times. Each of the newly created offspring is evaluated, and the parents and the offspring are added to the selection pool. The single best individual among the ![]() solutions in the pool survives and becomes the parent for the next iteration. Now we describe the
solutions in the pool survives and becomes the parent for the next iteration. Now we describe the ![]() strategy, which is a generalization of the
strategy, which is a generalization of the ![]() scheme, where
scheme, where ![]() parents generate
parents generate ![]() offspring through the recombination of
offspring through the recombination of ![]() individuals.
individuals.
 ES Algorithm
ES Algorithm
- Step 1. Initialize the population of size
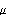 by randomly assigning locations
by randomly assigning locations  and strategy parameters
and strategy parameters  , where
, where  and
and  .
. - Step 2. Using the recombination operator
 , generate
, generate  offspring by randomly selecting and recombining
offspring by randomly selecting and recombining  individuals from the pool of
individuals from the pool of  parents.
parents. - Step 3. Mutations.
- Step 4. Evaluate the fitness
 of all individuals in
of all individuals in  .
. - Step 5. Select the
 best individuals to serve as parents for the next generation.
best individuals to serve as parents for the next generation. - Step 6. If the termination criterion is met:
2.2. Basic Particle Swarm Optimization
As the bPSO we use the original PSO version introduced by Eberhart and Kennedy [6]. Inspired by both social behavior and bird flocking patterns, the particles “fly” through the solution space and tend to land on better solutions.
The search is performed by a population of particles ![]() ; each has a location vector
; each has a location vector ![]() that represents a potential solution in a
that represents a potential solution in a ![]() -dimensional search space. Each particle
-dimensional search space. Each particle ![]() also keeps track of its velocity vector
also keeps track of its velocity vector ![]() that determines the direction and how far the particle moves in the next iteration. The fitness of a particle is determined by an evaluation function
that determines the direction and how far the particle moves in the next iteration. The fitness of a particle is determined by an evaluation function ![]() . Particles move through the search space in discrete time steps. In order to provide a balance between local (with a higher tendency to converge to a solution in close proximity) and global search (looking for overall good solutions), an inertia weight
. Particles move through the search space in discrete time steps. In order to provide a balance between local (with a higher tendency to converge to a solution in close proximity) and global search (looking for overall good solutions), an inertia weight ![]() was suggested by Shi and Eberhart [11, 12]. In the bPSO algorithm this term is set to the constant value
was suggested by Shi and Eberhart [11, 12]. In the bPSO algorithm this term is set to the constant value ![]() .
.
bPSO Algorithm
- Step 1. Initialize particle population of
 particles by stochastically assigning locations
particles by stochastically assigning locations  and velocities
and velocities  .
. - Step 2. Evaluate the fitness of all particles:
 .
. - Step 3. Keep track of the locations where each individual had its highest fitness so far:
- Step 4. Keep track of the position with the global best fitness:
- Step 5. Modify the particle velocities based on the previous best and global best positions:
- Step 6. Update the particle locations:
- Step 7. If the termination criterion is met:
2.3. Random Particle Swarm Optimization
Previous work with PSOs suggests that the so-called inertia weight ![]() should be annealed (dPSO) over time in order to obtain better results [13]. This time-decreasing inertia weight facilitates a global search at the beginning and the later small inertia weight fine-tunes the search space. Since the annealed value is dependent on time, the number of iterations must be known in advance. However, in most real-world scenarios, like the soccer-kick optimization [2], it is extremely difficult to know the number of necessary iteration steps in advance. The “random” rPSO version tries to alleviate this problem by assigning a random number to
should be annealed (dPSO) over time in order to obtain better results [13]. This time-decreasing inertia weight facilitates a global search at the beginning and the later small inertia weight fine-tunes the search space. Since the annealed value is dependent on time, the number of iterations must be known in advance. However, in most real-world scenarios, like the soccer-kick optimization [2], it is extremely difficult to know the number of necessary iteration steps in advance. The “random” rPSO version tries to alleviate this problem by assigning a random number to ![]() (Step 5) in each iteration as follows:
(Step 5) in each iteration as follows:
When ![]() is a uniformly distributed random number in the interval
is a uniformly distributed random number in the interval ![]() ,
, ![]() is a uniformly distributed random number in the interval
is a uniformly distributed random number in the interval ![]() .
.
3. Benchmarks
According to the no free lunch theorem [14], it is difficult to identify a clearly superior search or optimization algorithm in any comparison. Therefore our purpose is not to show which one of the three algorithms outperforms the others in any particular case, but to find out which of these optimizers is better suited for specific optimization challenges. In particular, we also want to investigate whether an algorithm’s performance characteristics in two dimensions—where visualization and manual inspection are easiest and most accessible—transfer to higher dimensions. We evaluate the performance of the ![]() ES scheme and both versions of the particle swarm algorithms on a small set of numerical benchmark functions.
ES scheme and both versions of the particle swarm algorithms on a small set of numerical benchmark functions.
We use the five benchmark functions illustrated and described in more detail following. We explore each of these benchmark search spaces for dimensions ![]() ,
, ![]() , and
, and ![]() . The first three functions are unimodal, that is, with a single global optimum. The last two functions are multimodal, where the number of local maxima increases exponentially with the problem size [15]. In the following function descriptions,
. The first three functions are unimodal, that is, with a single global optimum. The last two functions are multimodal, where the number of local maxima increases exponentially with the problem size [15]. In the following function descriptions, ![]() denotes the location of the global optimum. In the function plots, the location of the global optimum is marked by a red sphere.
denotes the location of the global optimum. In the function plots, the location of the global optimum is marked by a red sphere.



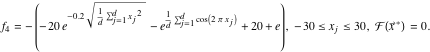


4. Experimental Setup
For each of the three optimizers ES, bPSO, and rPSO, we performed 20 experiments on each of the five test functions ![]() to
to ![]() for dimensions
for dimensions ![]() ,
, ![]() , and
, and ![]() . A different random number seed is used for each of the 20 runs. The initial individuals (including those for ES) are uniformly distributed throughout the search space. Each initial population generated for an ES experiment is also used for the bPSO and rPSO experiments. This ensures that all comparable runs start with the same initial distribution of individuals. The termination criterion for all runs was to stop when the maximum number of iterations
. A different random number seed is used for each of the 20 runs. The initial individuals (including those for ES) are uniformly distributed throughout the search space. Each initial population generated for an ES experiment is also used for the bPSO and rPSO experiments. This ensures that all comparable runs start with the same initial distribution of individuals. The termination criterion for all runs was to stop when the maximum number of iterations ![]() was reached. By that time all the optimizers had already reached their convergence phase (see Figure 2 later). The parameter settings for the three algorithms are described in Tables 1, 2, and 3.
was reached. By that time all the optimizers had already reached their convergence phase (see Figure 2 later). The parameter settings for the three algorithms are described in Tables 1, 2, and 3.


5. Discussion of the Results
The results of the evolution-based optimizer and the two particle swarm optimization techniques are briefly discussed in this section.
5.1. Phenotype Plots
Figure 1 gives an example of the population dynamics resulting from each of the three algorithms (ES in column 1, bPSO in column 2, and rPSO in column 3) applied over a certain number of iterations. The individuals are represented as dots where in each plot three iterations (red, blue, green) are depicted. In order to achieve a fair comparison, all three algorithms start from the same initial populations. The behavior of the individuals is seen at different iterations, making it easy to compare and contrast the movement of the individuals and study their convergence behavior. For example, ![]() is plotted at iterations 2 (red), 7 (blue), and 40 (green).
is plotted at iterations 2 (red), 7 (blue), and 40 (green).
In comparison to the ![]() ES scheme, we observe that the particle swarm individuals (both bPSO and rPSO) have higher exploration capabilities, search the solution space more thoroughly, and search in multiple directions. The ES individuals stay close to each other within a certain mutation radius. This is a typical effect of using this particular ES scheme. The individuals of the ES algorithm converge to a local optimum solution for functions
ES scheme, we observe that the particle swarm individuals (both bPSO and rPSO) have higher exploration capabilities, search the solution space more thoroughly, and search in multiple directions. The ES individuals stay close to each other within a certain mutation radius. This is a typical effect of using this particular ES scheme. The individuals of the ES algorithm converge to a local optimum solution for functions ![]() and
and ![]() . This also illustrates the fact that ES individuals exhibit strong local search behaviors, which in this case is mainly due to a relatively small mutation step size.
. This also illustrates the fact that ES individuals exhibit strong local search behaviors, which in this case is mainly due to a relatively small mutation step size.

Figure 1. Phenotypical plots for the two-dimensional versions of the benchmark functions ![]() to
to ![]() . From left to right the columns show snapshots of typical optimization runs for ES, bPSO, and rPSO. The populations of the first, second, and third snapshot at the iterations as indicated are represented as red, blue, and green spheres, respectively.
. From left to right the columns show snapshots of typical optimization runs for ES, bPSO, and rPSO. The populations of the first, second, and third snapshot at the iterations as indicated are represented as red, blue, and green spheres, respectively.
5.2. Convergence Plots
The convergence plots represent mean fitness values computed over all 20 runs for all 1500 iterations. This graph helps to demonstrate the convergence behavior of the individuals of a particular algorithm and also illustrates which of the three algorithms has the fastest fitness convergence speed. Figure 2 summarizes the results, which are shown for ![]() (column 1),
(column 1), ![]() (column 2), and
(column 2), and ![]() (column 3).
(column 3).
In almost all cases, the ES algorithm is comparable to rPSO. The bPSO algorithm turns out to be the slowest in terms of convergence speed for ![]() . However, as the number of dimensions increases (
. However, as the number of dimensions increases (![]() ), the convergence rate of ES decreases, and in 10 dimensions ES converges more slowly and toward lower fitness values.
), the convergence rate of ES decreases, and in 10 dimensions ES converges more slowly and toward lower fitness values.
All three algorithms show relatively steep ascents during the first 100 or 200 iterations. In general, rPSO tends to rapidly level off without making any further progress (for most of the test cases) during the rest of the simulation; that is, the swarm stagnates and no changes are observed in terms of finding a better fitness value. For instance, on ![]() , particle swarms stagnate and flatten out without any further improvements. However, the convergence rate of ES on the function
, particle swarms stagnate and flatten out without any further improvements. However, the convergence rate of ES on the function ![]() gradually slows down but does not completely level off, which indicates that if it is allowed to run longer it may discover better solutions.
gradually slows down but does not completely level off, which indicates that if it is allowed to run longer it may discover better solutions.
Another observation made for function ![]() in two dimensions is that rPSO has the slowest convergence rate. This is in line with the results of the phenotype plot (Figure 1), where the particles do not converge to one location.
in two dimensions is that rPSO has the slowest convergence rate. This is in line with the results of the phenotype plot (Figure 1), where the particles do not converge to one location.

Figure 2. Fitness plots of the benchmark functions ![]() to
to ![]() . From left to right the columns illustrate the convergence behaviors of the three algorithms (ES: blue, rPSO: orange, bPSO: green) in 2, 4, and 10 dimensions.
. From left to right the columns illustrate the convergence behaviors of the three algorithms (ES: blue, rPSO: orange, bPSO: green) in 2, 4, and 10 dimensions.
5.3. Success Plots
The best fitness value obtained at the end of each run is illustrated in Figure 3. For each function the fitness values of all 20 runs are plotted in ascending order from left to right. Therefore, each graph displays the success rate of each algorithm on a particular function. The best (right-most point), worst (left-most point), and mean fitness can also be easily derived from these graphs.
In all 20 runs, the ES algorithm finds worse solutions than both PSO algorithms for ![]() , 4, and 10, as shown by the left-most blue point in Figure 3. This is in line with the results of the phenotype plots (Figure 1), especially for the multimodal functions
, 4, and 10, as shown by the left-most blue point in Figure 3. This is in line with the results of the phenotype plots (Figure 1), especially for the multimodal functions ![]() and
and ![]() , where the ES individuals are unsuccessful in finding the global optimum. The higher exploration capabilities of the PSO algorithms seem to facilitate the discovery of better solutions in comparison to the local ES scheme.
, where the ES individuals are unsuccessful in finding the global optimum. The higher exploration capabilities of the PSO algorithms seem to facilitate the discovery of better solutions in comparison to the local ES scheme.
In the phenotype plots (Figure 1) it can also be observed that the bPSO particles do not converge to one solution only. However, there is always at least one particle that finds the global optimum. Therefore, the bPSO and rPSO algorithms are comparable in terms of finding the best solution, as illustrated by the right-most green and orange points in Figure 3.

Figure 3. Success ratio plots of the benchmark functions ![]() to
to ![]() . From left to right the columns show the best fitness value obtained at the end of each run of the three algorithms (ES: blue, rPSO: orange, bPSO: green) in 2, 4, and 10 dimensions.
. From left to right the columns show the best fitness value obtained at the end of each run of the three algorithms (ES: blue, rPSO: orange, bPSO: green) in 2, 4, and 10 dimensions.
5.4. Parameter Range Plots
For the following analysis we only look at algorithm performance on the 10-dimensional benchmarks. In Figure 4 we visualize changes during the course of the evolutionary search for each variable range. For example, in the first row of Figure 4 the vertical bars represent the range for each of the 10 variables, over all iterations, limited to all those solutions that have a fitness of at least ![]() . Here the highest fitness is zero. Knowing how the value ranges change is important, especially when exploring real-world optimization problems, since it can provide insights on whether the fitness function is sensitive with respect to a particular variable or not. Figure 4 also clearly visualizes the stage of convergence in each dimension.
. Here the highest fitness is zero. Knowing how the value ranges change is important, especially when exploring real-world optimization problems, since it can provide insights on whether the fitness function is sensitive with respect to a particular variable or not. Figure 4 also clearly visualizes the stage of convergence in each dimension.
Figure 4 shows that the ES algorithm maintains a high parameter range in comparison to the particle swarm algorithms. This is in line with the results of Figure 2, where ES has the slowest convergence speed for ![]() . Evolution strategies seem to consistently keep wider parameter ranges. Both particle swarm algorithms show comparable ranges.
. Evolution strategies seem to consistently keep wider parameter ranges. Both particle swarm algorithms show comparable ranges.

Figure 4. Parameter range plots for the 10-dimensional versions of the benchmark functions ![]() to
to ![]() . From left to right the columns show the parameter ranges for each of the 10 dimensions over a certain threshold value (
. From left to right the columns show the parameter ranges for each of the 10 dimensions over a certain threshold value (![]() ) for ES, bPSO, and rPSO.
) for ES, bPSO, and rPSO.
6. “Evolutionary Swarms” ![webAdjustmentBox[Mathematica, BoxMargins -> {{-0.175, 0}, {0, 0}}] webAdjustmentBox[Mathematica, BoxMargins -> {{-0.175, 0}, {0, 0}}]](https://content.wolfram.com/sites/19/2011/01/Khemka_Image_10.gif) Site
Site
The comparisons conducted provide knowledge and insights about the algorithms, while illustrating the movement of the individuals in the solution space (Figure 1). They point out various characteristics such as the convergence speed and the success an algorithm has in finding good solutions. The strengths and weaknesses of the algorithms are exploited. For example, PSO algorithms have a higher exploration rate, whereas the ![]() scheme of ES depicts a local search scheme.
scheme of ES depicts a local search scheme.
We created a set of notebooks to compare the ![]() scheme of ES and both of the particle swarm optimizers on the benchmark functions (
scheme of ES and both of the particle swarm optimizers on the benchmark functions (![]() to
to ![]() ). We then converted those notebooks to webMathematica. This interactive site provides a hands-on tutorial and an experimental environment through a selection of 10 benchmark functions along with visualization tools.
). We then converted those notebooks to webMathematica. This interactive site provides a hands-on tutorial and an experimental environment through a selection of 10 benchmark functions along with visualization tools.
This site currently includes three variants of particle swarms: basic (bPSO), random (rPSO), and those with decreasing inertia weight (dPSO). We also implemented both of the simple ES schemes: the ![]() scheme and the
scheme and the ![]() scheme as well as the generalized evolution strategies
scheme as well as the generalized evolution strategies ![]() and
and ![]() as described in Section 2.
as described in Section 2.
As it is in general difficult to know the settings of various parameters, we provide suggestions for different settings. This will help users to gain further knowledge regarding these optimizers.
The website can be accessed at www.swarm-design.org.
7. Conclusion
The best way to understand and use evolution- and swarm-based algorithm heuristics is through practical experience, which can be gained most efficiently on smaller-scale problems. The Evolutionary & Swarm Optimization website that we developed will be merged with the collection of notebooks from the Evolvica package [16]. This database of notebooks and the swarm algorithms provides an experimental and inquiry platform for introducing evolutionary and swarm-based optimization techniques to those who wish to further their knowledge of evolutionary computation. Making these notebooks available through a ![]() site means that anyone with access to the newly built web pages will have instant access to a wide range of optimization algorithms.
site means that anyone with access to the newly built web pages will have instant access to a wide range of optimization algorithms.
References
| [1] | C. Jacob and N. Khemka, “Particle Swarm Optimization: An Exploration Kit for Evolutionary Optimization,” in New Ideas in Symbolic Computation: Proceedings of the Sixth International Mathematica Symposium (IMS’04), Banff, Alberta, Canada (P. Mitic, C. Jacob, and J. Carne, eds.), Hampshire, UK: Positive Corporation Ltd., 2004. library.wolfram.com/infocenter/Conferences/6039. |
| [2] | N. Khemka, C. Jacob, and G. Cole, “Making Soccer Kicks Better: A Study in Particle Swarm Optimization and Evolution Strategies,” in Proceedings of the IEEE Congress on Evolutionary Computation (CEC’05), Edinburgh, U.K., New York: IEEE, 2005. |
| [3] | D. Goldberg, Genetic Algorithms in Search, Optimization, and Machine Learning, Reading, MA: Addison-Wesley Publishing Company, 1989. |
| [4] | J. R. Koza, Genetic Programming: On the Programming of Computers by Means of Natural Selection, Cambridge, MA: MIT Press, 1992. |
| [5] | I. Rechenberg, “Evolution Strategies: Nature’s Way of Optimization,” Optimization Methods and Applications, Possibilities and Limitations (H. W. Bergmann, ed.), Lecture Notes in Computer Science, 47, Berlin: Springer, 1989 pp. 106-126. |
| [6] | J. Kennedy and R. C. Eberhart, “Particle Swarm Optimization,” in Proceedings of the IEEE International Conference on Neural Networks, Perth, WA, Australia, New York: IEEE, 1995 pp. 1942-1948. |
| [7] | G. Cole and K. Gerristen, Influence of Mass Distribution in the Shoe and Plate Stiffness on Ball Velocity During a Soccer Kick, Herzogenaurach, Germany: Adidas-Salomon AG, 2002. |
| [8] | G. Cole and K. Gerristen, Optimal Mass Distribution and Plate Stiffness of Football Shoes, Herzogenaurach, Germany: Adidas-Salomon AG, 2002. |
| [9] | G. Cole and K. Gerristen, Influence of Medio-Lateral Mass Distribution in a Soccer Shoe on the Deflection of the Ankle and Subtabular Joints during Off-Centre Kicks, Herzogenaurach, Germany: Adidas-Salomon AG, 2003. |
| [10] | N. Khemka, “Comparing PSO and ES: Benchmarks and Applications,” M.Sc. Thesis, University of Calgary, Calgary, Alberta, Canada, 2005. |
| [11] | Y. Shi and R. C. Eberhart, “Parameter Selection in Particle Swarm Optimization,” in Evolutionary Programming VII: Proceedings of the Seventh International Conference on Evolutionary Programming, San Diego, CA (V. W. Porto, N. Saravanan, D. E. Waagen, and A. E. Eiben, eds.), Lecture Notes in Computer Science, 1447, London: Springer-Verlag, 1998 pp. 591-600. |
| [12] | Y. Shi and R. C. Eberhart, “A Modified Particle Swarm Optimizer,” in Proceedings of the IEEE Congress on Evolutionary Computation (CEC’98), Anchorage, AK, New York: IEEE, 1998 pp. 69-73. |
| [13] | R. C. Eberhart and Y. Shi, “Comparison between Genetic Algorithms and Particle Swarm Optimization,” in Evolutionary Programming VII: Proceedings of the Seventh International Conference on Evolutionary Programming, San Diego, CA (V. W. Porto, N. Saravanan, D. E. Waagen, and A. E. Eiben, eds.), Lecture Notes in Computer Science, 1447, London: Springer-Verlag, 1998 pp. 611-616. |
| [14] | D. H. Wolpert and W. G. Macready, “No Free Lunch Theorems for Search,” SFI Working Paper # 95-02-010, Santa Fe, NM: Santa Fe Institute, 1995. |
| [15] | H-P. Schwefel, Evolution and Optimum Seeking, New York: John Wiley & Sons, 1995. |
| [16] | C. Jacob, Illustrating Evolutionary Computation with Mathematica (The Morgan Kaufmann Series in Artificial Intelligence), San Francisco, CA: Morgan Kaufmann Publishers, 2001. |
| Namrata Khemka, and Christian Jacob, “Exploratory Toolkit for Evolutionary and Swarm-Based Optimization,” The Mathematica Journal, 2010. dx.doi.org/doi:10.3888/tmj.11.3-5. | |
About the Authors
Namrata Khemka received her Ph.D. in computer science from the University of Calgary in 2009. She received her M.Sc. in 2005 and B.Sc. in 2003. Her interests lie in data visualization, optimization techniques, and swarm- and agent-based modeling.
Christian Jacob received his Ph.D. in computer science from the University of Erlangen-Nuremberg in Erlangen, Germany. He is currently an associate professor in the Department of Computer Science (Faculty of Science) and the Department of Biochemistry & Molecular Biology (Faculty of Medicine) at the University of Calgary. Jacob’s research interests are in evolutionary computing, emergent phenomena, and swarm intelligence, with applications in civil engineering, biological modeling, medical sciences, computational creativity, and art.
Namrata Khemka
Christian Jacob
University of Calgary, Calgary, AB, Canada, T2N1N4
namrata.khemka@gmail.com
jacob@cpsc.ucalgary.ca